Research concept
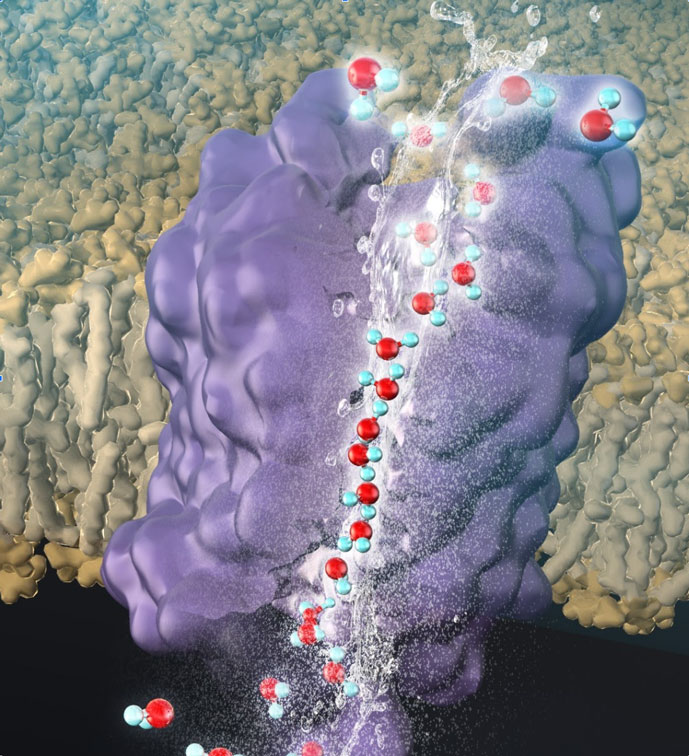
Understanding of the principles of protein function on the basis of the molecular structure
Proteins consist of only 20 types of amino acids, while they show large variety in their functions, e.g., redox activity, transporter, sensor, and antibodies. Protein functions can be fully explained by the molecular structure even if the functions are seemingly complicated. To clarify a relationship between functions and structures of proteins, we analyze protein structures at an atomic level and calculate physical or chemical constants on the basis of theoretical chemistry. “Just computing molecules” is not in our interest. Our mission is to uncover new but simple principles essential to the protein science. For example, we are trying to clarify the reaction mechanisms of natural photosynthetic proteins, e.g., oxygen evolution, electron transfer, and proton transfer reactions. We also develop new tools for analysis of protein function.

Theme 1:Light energy conversion
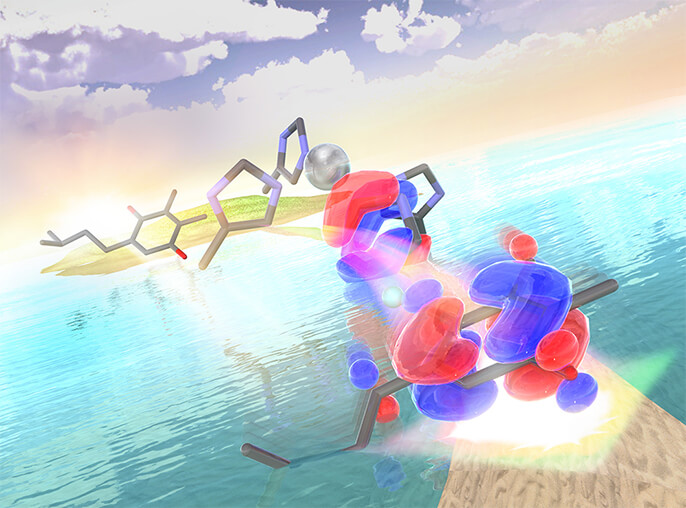
In plant photosynthesis, light energy is converted into a flow of electrons and H+ (proton). The photosynthetic reaction is catalyzed by various proteins that receive light. The retina of our eyes contains photorecepter proteins that receives light and emits a signal as a optical sensor of a digital camera. We use all tools such as molecular chemistry, physical chemistry, photochemistry, inorganic chemistry, and complex chemistry to understand the reaction mechanism of these photoactive proteins.
Theme 2:Enzyme, catalyst, and drug design
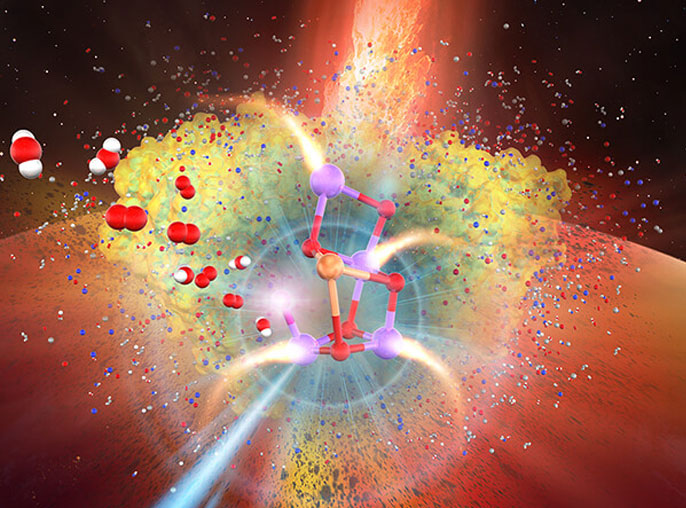
Enzymes are proteins that catalyze physiological reactions in our body. Inhibitor (drug) molecules bind to enzymes and inhibits enzymatic reactions. We are investigating how the enzymatic reaction occurs and what inhibitor molecule acts effectively as a drug.
Theme 3:Development of new method in theoretical chemistry

We conduct research using theoretical and computational methods such as quantum chemistry, electrostatic calculations, molecular dynamics simulations, and machine learning. By combining these methods, we are also developing a method for calculating pKa, redox potential, absorption wavelength of molecules (in the protein environment).